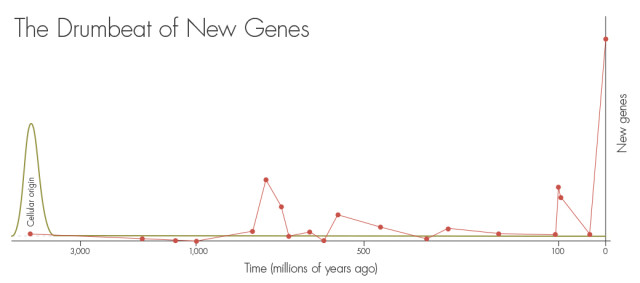
How new genes arise from junk DNA

Genes, like people, have families — lineages that stretch back through time, all the way to a founding member. That ancestor multiplied and spread, morphing a bit with each new iteration.
For most of the last 40 years, scientists thought that this was the primary way new genes were born — they simply arose from copies of existing genes. The old version went on doing its job, and the new copy became free to evolve novel functions.
Certain genes, however, seem to defy that origin story. They have no known relatives, and they bear no resemblance to any other gene. They’re the molecular equivalent of a mysterious beast discovered in the depths of a remote rainforest, a biological enigma seemingly unrelated to anything else on earth.
The mystery of where these orphan genes came from has puzzled scientists for decades. But in the past few years, a once-heretical explanation has quickly gained momentum — that many of these orphans arose out of so-called junk DNA, or non-coding DNA, the mysterious stretches of DNA between genes. “Genetic function somehow springs into existence,” said David Begun, a biologist at the University of California, Davis.
This metamorphosis was once considered to be impossible, but a growing number of examples in organisms ranging from yeast and flies to mice and humans has convinced most of the field that these de novo genes exist. Some scientists say they may even be common. Just last month, research presented at the Society for Molecular Biology and Evolution in Vienna identified 600 potentially new human genes. “The existence of de novo genes was supposed to be a rare thing,” said Mar Albà, an evolutionary biologist at the Hospital del Mar Research Institute in Barcelona, who presented the research. “But people have started seeing it more and more.”
Researchers are beginning to understand that de novo genes seem to make up a significant part of the genome, yet scientists have little idea of how many there are or what they do. What’s more, mutations in these genes can trigger catastrophic failures. “It seems like these novel genes are often the most important ones,” said Erich Bornberg-Bauer, a bioinformatician at the University of Münster in Germany.
The Orphan Chase
The standard gene duplication model explains many of the thousands of known gene families, but it has limitations. It implies that most gene innovation would have occurred very early in life’s history. According to this model, the earliest biological molecules 3.5 billion years ago would have created a set of genetic building blocks. Each new iteration of life would then be limited to tweaking those building blocks.
Yet if life’s toolkit is so limited, how could evolution generate the vast menagerie we see on Earth today? “If new parts only come from old parts, we would not be able to explain fundamental changes in development,” Bornberg-Bauer said.
The first evidence that a strict duplication model might not suffice came in the 1990s, when DNA sequencing technologies took hold. Researchers analyzing the yeast genome found that a third of the organism’s genes had no similarity to known genes in other organisms. At the time, many scientists assumed that these orphans belonged to families that just hadn’t been discovered yet. But that assumption hasn’t proven true. Over the last decade, scientists sequenced DNA from thousands of diverse organisms, yet many orphan genes still defy classification. Their origins remain a mystery.
In 2006, Begun found some of the first evidence that genes could indeed pop into existence from noncoding DNA. He compared gene sequences from the standard laboratory fruit fly, Drosophila melanogaster, with other closely related fruit fly species. The different flies share the vast majority of their genomes. But Begun and collaborators found several genes that were present in only one or two species and not others, suggesting that these genes weren’t the progeny of existing ancestors. Begun proposed instead that random sequences of junk DNA in the fruit fly genome could mutate into functioning genes.
Yet creating a gene from a random DNA sequence appears as likely as dumping a jar of Scrabble tiles onto the floor and expecting the letters to spell out a coherent sentence. The junk DNA must accumulate mutations that allow it to be read by the cell or converted into RNA, as well as regulatory components that signify when and where the gene should be active. And like a sentence, the gene must have a beginning and an end — short codes that signal its start and end.
In addition, the RNA or protein produced by the gene must be useful. Newly born genes could prove toxic, producing harmful proteins like those that clump together in the brains of Alzheimer’s patients. “Proteins have a strong tendency to misfold and cause havoc,” said Joanna Masel, a biologist at the University of Arizona in Tucson. “It’s hard to see how to get a new protein out of random sequence when you expect random sequences to cause so much trouble.” Masel is studying ways that evolution might work around this problem.
Another challenge for Begun’s hypothesis was that it’s very difficult to distinguish a true de novo gene from one that has changed drastically from its ancestors. (The difficulty of identifying true de novo genes remains a source of contention in the field.)
Ten years ago, Diethard Tautz, a biologist at the Max Planck Institute for Evolutionary Biology, was one of many researchers who were skeptical of Begun’s idea. Tautz had found alternative explanations for orphan genes. Some mystery genes had evolved very quickly, rendering their ancestry unrecognizable. Other genes were created by reshuffling fragments of existing genes.
Then his team came across the Pldi gene, which they named after the German soccer player Lukas Podolski. The sequence is present in mice, rats and humans. In the latter two species, it remains silent, which means it’s not converted into RNA or protein. The DNA is active or transcribed into RNA only in mice, where it appears to be important — mice without it have slower sperm and smaller testicles.
The researchers were able to trace the series of mutations that converted the silent piece of noncoding DNA into an active gene. That work showed that the new gene is truly de novo and ruled out the alternative — that it belonged to an existing gene family and simply evolved beyond recognition. “That’s when I thought, OK, it must be possible,” Tautz said.
A Wave of New Genes
Scientists have now catalogued a number of clear examples of de novo genes: A gene in yeast that determines whether it will reproduce sexually or asexually, a gene in flies and other two-winged insects that became essential for flight, and some genes found only in humans whose function remains tantalizingly unclear.
The Odds of Becoming a Gene
Scientists are testing computational approaches to determine how often random DNA sequences can be mutated into functional genes. Victor Luria, a researcher at Harvard, created a model using common estimates of the rates of mutation, recombination (another way of mixing up DNA) and natural selection. After subjecting a stretch of DNA as long as the human genome to mutation and recombination for 100 million generations, some random stretches of DNA evolved into active genes. If he were to add in natural selection, a genome of that size could generate hundreds or even thousands of new genes.
At the Society for Molecular Biology and Evolution conference last month, Albà and collaborators identified hundreds of putative de novo genes in humans and chimps — ten-fold more than previous studies — using powerful new techniques for analyzing RNA. Of the 600 human-specific genes that Albà’s team found, 80 percent are entirely new, having never been identified before.
Unfortunately, deciphering the function of de novo genes is far more difficult than identifying them. But at least some of them aren’t doing the genetic equivalent of twiddling their thumbs. Evidence suggests that a portion of de novo genes quickly become essential. About 20 percent of new genes in fruit flies appear to be required for survival. And many others show signs of natural selection, evidence that they are doing something useful for the organism.
In humans, at least one de novo gene is active in the brain, leading some scientists to speculate such genes may have helped drive the brain’s evolution. Others are linked to cancer when mutated, suggesting they have an important function in the cell. “The fact that being misregulated can have such devastating consequences implies that the normal function is important or powerful,” said Aoife McLysaght, a geneticist at Trinity College in Dublin who identified the first human de novo genes.
Promiscuous Proteins
De novo genes are also part of a larger shift, a change in our conception of what proteins look like and how they work. De novo genes are often short, and they produce small proteins. Rather than folding into a precise structure — the conventional notion of how a protein behaves — de novo proteins have a more disordered architecture. That makes them a bit floppy, allowing the protein to bind to a broader array of molecules. In biochemistry parlance, these young proteins are promiscuous.
Scientists don’t yet know a lot about how these shorter proteins behave, largely because standard screening technologies tend to ignore them. Most methods for detecting genes and their corresponding proteins pick out long sequences with some similarity to existing genes. “It’s easy to miss these,” Begun said.
That’s starting to change. As scientists recognize the importance of shorter proteins, they are implementing new gene discovery technologies. As a result, the number of de novo genes might explode. “We don’t know what things shorter genes do,” Masel said. “We have a lot to learn about their role in biology.”
Scientists also want to understand how de novo genes get incorporated into the complex network of reactions that drive the cell, a particularly puzzling problem. It’s as if a bicycle spontaneously grew a new part and rapidly incorporated it into its machinery, even though the bike was working fine without it. “The question is fascinating but completely unknown,” Begun said.
A human-specific gene called ESRG illustrates this mystery particularly well. Some of the sequence is found in monkeys and other primates. But it is only active in humans, where it is essential for maintaining the earliest embryonic stem cells. And yet monkeys and chimps are perfectly good at making embryonic stem cells without it. “It’s a human-specific gene performing a function that must predate the gene, because other organisms have these stem cells as well,” McLysaght said.
“How does novel gene become functional? How does it get incorporated into actual cellular processes?” McLysaght said. “To me, that’s the most important question at the moment.”
This article is published in collaboration with Quanta Magazine. Publication does not imply endorsement of views by the World Economic Forum.
To keep up with the Agenda subscribe to our weekly newsletter.
Author: Emily Singer is a senior writer and contributing editor at Quanta Magazine covering the life sciences.
Image: A DNA double helix is seen in an undated artist’s illustration released by the National Human Genome Research Institute to Reuters on May 15, 2012. REUTERS/National Human Genome Research Institute/Handout.
Don't miss any update on this topic
Create a free account and access your personalized content collection with our latest publications and analyses.
License and Republishing
World Economic Forum articles may be republished in accordance with the Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International Public License, and in accordance with our Terms of Use.
The views expressed in this article are those of the author alone and not the World Economic Forum.
Stay up to date:
Emerging Technologies
Forum Stories newsletter
Bringing you weekly curated insights and analysis on the global issues that matter.
More on Emerging TechnologiesSee all
Sercan Esen
May 27, 2026